- Volume 60 , Number 3
- Page: 482–3
Polymerase chain reaction applied to biopsies f rom paucibacillary leprosy
This department is for the publication of informal communications that are of interest because they are informative and for the discussion of controversial matters. The mandate of this JORNAL is to disseminate information relating to leprosy in particular and also other mycobacterial diseases. Dissident comment or interpretation on published research is of course valid, but personality attacks on individuals would seem unnecessary. Political comments, valid or not, also are unwelcome. They might result in interference with the distribution of the JOURNAL and thus interfere with its prime purpose.
To the Editor:
In a previous study (2), we applied the polymerase chain reaction (PCR) to mouse foot pad harvests and human skin biopsies from multibacillary (MB) patients. One of the practical applications of the PCR could be the documentation of paucibacillary (PB) leprosy, which is the subject of the present study.
MATERIALS AND METHODS
Ten 5-mm punch skin biopsies were taken from 10 different patients with the clinical diagnosis of paucibacillary (9 cases) or indeterminate leprosy (1 case) at the Institut Marehoux, Bamako, Mali. The biopsies were divided longitudinally into halves; one was fixed in 10% Formalin for histologic diagnosis, the other was placed in a dry container on ice. The two series of biopsies were sent by air to the Leprosy Laboratory of the Institute for Tropical Medicine, Antwerp, Belgium, where they arrived 48 hr later.
The fresh biopsies were weighed, minced with HCl-cleaned sterilized scissors, suspended in 0.5-ml distilled water, and stored at 20ºC until PCR processing. The Formalin-fixed specimens were embedded in paraffin, and the sections were stained with the Triff technique (1).
The sample preparation for the PCR consisted of five cycles of freeze-thawing; the PCR was performed as described previously(2). The reaction was done simultaneously in duplicate on each suspension: one tube with, and a second tube without, an internal control. Furthermore, between each couple of tubes a blank containing distilled water instead of a biopsy suspension was introduced as the internal control. This tube was subjected to exactly the same manipulations as were the biopsies. When PCR inhibitors were detected in the tube containing the internal control, the reaction was repeated with a 1/10 dilution of the sample.
RESULTS AND DISCUSSION
On histopathological examination, two cases were diagnosed as tuberculoid (TT) and seven as borderline tuberculoid (BT) leprosy. In the possible indeterminate case clinically only some perivascular lymphocytic infiltration was seen, and no definite diagnosis of leprosy could be made. Acidfast bacilli were not found in any of three sections of each case.
As shown in The Table, 3 specimens produced a positive result, 1 was negative and inhibitors were present in 6. When the latter was diluted 10-1, one specimen still contained inhibitors while the five others gave a negative reaction.
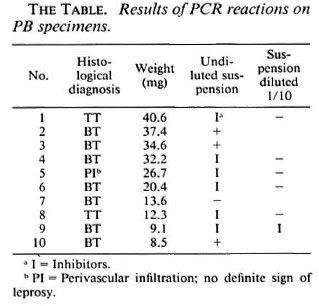
It is striking that, with the exception of biopsy number 1, 2 of the 3 positive results were obtained among the three heaviest biopsies. It seems that a positive result can only exceptionally be obtained in biopsies weighing 30 mg or less.
These results confirm our earlier predictions based on the PCR results from MB leprosy biopsies (2), that the technique as presently performed can only detect Mycobacterium leprae DNA in some PB cases, probably those containing the higher number of M. leprae still compatible with PB leprosy.
- Stefaan R. Pattyn, M.D.
Professor of Medical Microbiology
University of Antwerp and Institute for Tropical Medicine
Nationalestraat 155
B-2000 Antwerp, Belgium
- Pierre Jamet, M.D.
Institut Marchoux
Bamako, Mali
- Veerle Raes
Laboratory Technician
Institute for Tropical Medicine
Antwerp, Belgium
REFERENCES
1. PATTYN, S. R., DOCKX, P. and CAP, J. A. La Lépre. Paris: Masson, 1981.
2. PATTYN, S. R., URSI, D., IEVEN, M. and RAES, V.Polymerase chain reaction, amplifying the DNA coding for species-specific rRNA of Mycobacterium leprae. Int. J. Lepr. 60(1992)234-243.