- Volume 62 , Number 1
- Page: 139–40
Detection of Mycobacterium leprae in tissue and blood by polymerase chain reaction
To the Editor:
Leprosy is a chronic, systemic, infectious disease caused by Mycobacterium leprae . A new diagnostic method which is more sensitive and specific for the detection of M. leprae is required which might allow diagnosis of leprosy at a very early stage. Recently, specific DNA probes have been developed, and several investigators have studied the use of the polymerase chain reaction (PCR) to detect M. leprae (1-4). Our present study applied PCR on sections of paraffin-embedded, frozen biopsy samples and peripheral blood from leprosy patients. Specimens were taken from 35 leprosy patients (19 males and 16 females between 14 and 73 years of age). Genomic DNA was isolated from paraffin-embedded or frozen tissues and peripheral blood, as described elsewhere. A set of primers, S13 (5'-CTCCACCTGGACCGGCGAT-3') and S62 (5'-GACTAGCCTGCCAAGTCG-3'), was selected on the basis of the nucleotide sequence positioned 530 bp of a gene encoding 36-kDa antigen of M. leprae (2). DNA samples were first heated at 94ºC for 10 min to denature the DNA and then subjected to 32 amplification cycles. For direct gel analysis 10 µ l of the reaction mixture was subjected to electrophoresis on an 1.2% agarose gel for 45 min at 50 volts, and DNA was visualized as UV fluorescence (320 nm) after staining with ethidium bromide. For Southern blot analysis, DNA was separated by electrophoresis and transferred to nitrocellulose (NC) paper after treatment with 1.5 M NaCl, 0.5 N NaOH (denaturation solution) for 20 min then the filter was neutralized with 1.5 M NaCl, 1 M Tris-HCl (neutralizing solution) for 20 min at room temperature. Thereafter the DNA on the gel was transferred to NC paper for 20 hr at room temperature, and then baked for 60 min at 80ºC. DNA samples were hybridized with digoxigenin-labeled M. leprae cDNA probe in hybridization solution. The filters were washed twice with 2 x SSC (1 x SSC is 0.15 M NaCl plus 0.015 M sodium citrate), 0.1% sodium dodecyl sulfate, incubated in a blocking buffer for 30 min, and treated with alkaline phosphate for 1 hr for the color reaction.
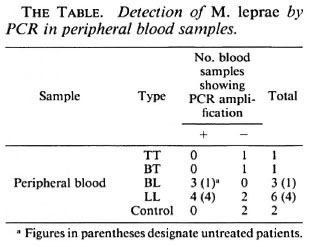
In frozen and paraffin-embedded tissues of leprosy patients, relatively high detection rates of amplified PCR products were achieved by using direct gel analysis as well as Southern blot hybridization (The Figure). In the peripheral blood of five untreated leprosy patients and two patients suffering from erythema nodosum leprosum (ENL), positive amplified PCR products were detected, indicating that M. leprae exist for a relatively long period in the peripheral blood of active cases (The Table).
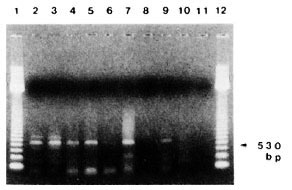
The figure. PCR detection of M. leprae DNA inbiopsy samples from leprosy patients. DNA from: purified M. leprae (lane 2), frozen sections from LL (lanes3-5) and BL (lane 6), paraffin-fixed sections from LL(lanes 7, 8), BT (lane 9) and TT (lane 10), and no DNA(negative control, lane 11). Lanes 1 and 12 containedmolecular size markers.
This study assessed the applicability of PCR coupled with DNA hybridization analysis for the detection of small numbers of M. leprae . Two set of primers, S 13 and S62, were selected on the basis of established nucleotide sequences of the 36-kDa genes to specifically detect M. leprae through amplification of characteristic DNA sequences. When we applied PCR on paraffinembedded and frozen biopsy tissues from leprosy patients, the frozen section results were better than those of the paraffin-embedded sections. This was especially so in the paraffin-embedded section from a bacterial index-negative treated patient who was PCR positive. A similar result has been reported by de Wit, et al. (1). When we applied PCR on peripheral blood, positive amplified PCR products were detected in five untreated leprosy patients and two patients with ENL, suggesting that M. leprae exist for a relatively long period in the peripheral blood of active cases. In this report apparent identification of M. leprae DNA in the tissue and blood by PCR shows that this method can be an additional tool for the diagnosis of suspected cases of early leprosy and for epidemiologic study.
- Kyu Suk Lee, M.D., Ph.D.
Oh Kwang Youl, M.D.
Ryoo Young Wok, M.D.
Department of Dermatology
- Min Ho Suh, M.D., Ph.D.
Department of Microbiology
Keimyung University School of Medicine
194 Dongsan-dong
Taegu, Korea
Acknowledgment. This investigation was supported by the Special Research Fund (1993) of the Dongsan Medical Center.
REFERENCES
1. DE WIT M. Y. L., FABER, W. R., KRIEG, S. R., DOUGLAS, J. T., LUCAS, S. B., MONTREEWASUWAT, N., PATTYN, S. R., HUSSAIN, R., PONNIGHAUS, J. M., HARTSKEERL, R. A. and KLATSER, P. R. Multiplication of a polymerase chain reaction for the detection of M. leprae in skin tissue. J. Clin. Microbiol. 29(1991)906-910.
2. HARTSKEERL, R. A., DE WIT, M. Y. L. and KLATSER, P. R. Polymerase chain reaction for the detection of Mycobacterium leprae . J. Gen. Microbiol. 135(1989)2357-2364.
3. PLIKAYTIS, B. B., GELBER, R. H. and SHINNICK, T. M. Rapid and sensitive detection of Mycobacterium leprae using a nested-primer gene amplification assay. J. Clin. Microbiol. 28(1990)1913-1917.
4. WILLIAMS, D. L., GILLIS, T. P., BOOTH, R. J., LOOKER, D. and WATSON, J. D. The use of a specific DNA probe and polymerase chain reaction for the detection of Mycobacterium leprae . J. Infect. Dis. 162(1990)193-200.